Scientific Papers
MycoKey aims to unlock new knowledge and to valorize existing knowledge and data by rapid dissemination to research partners and stakeholders in the chain.
In the MycoKey programme all scientific peer reviewed publications are available through open access. Research data will be deposited in public data repositories for (re)-analysis exploitation and dissemination free of charge.
In this page you can find publications describing the scientific results arising from project’s activities.
Evolution and Diversity of Biosynthetic Gene Clusters in Fusarium
- google+
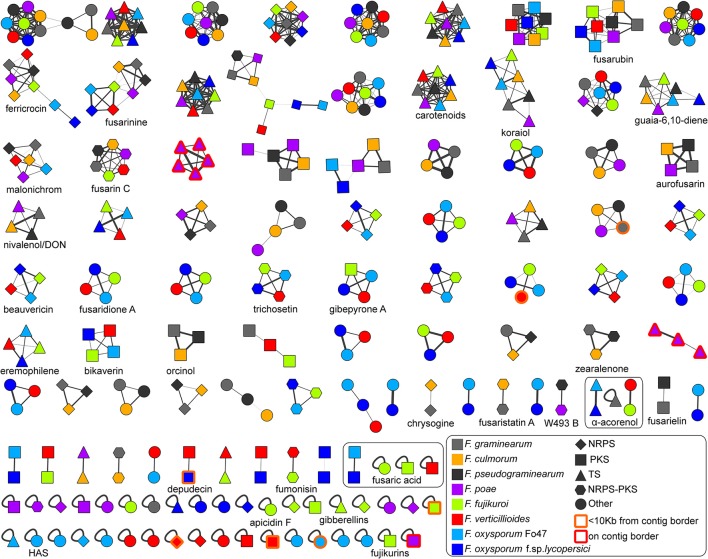
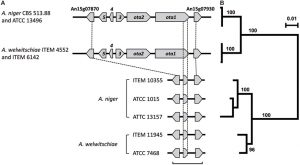
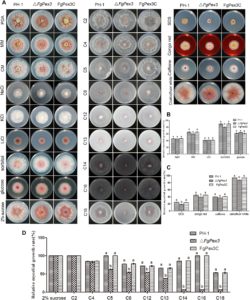
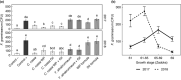
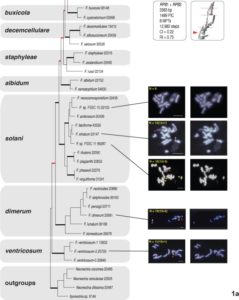
Plant pathogenic fungi in the Fusarium genus cause severe damage to crops, resulting in great financial losses and health hazards. Specialized metabolites synthesized by these fungi are known to play key roles in the infection process, and to provide survival advantages inside and outside the host. However, systematic studies of the evolution of specialized metabolite-coding potential across Fusarium have been scarce. Here, we apply a combination of bioinformatic approaches to identify biosynthetic gene clusters (BGCs) across publicly available genomes from Fusarium, to group them into annotated families and to study gain/loss events of BGC families throughout the history of the genus. Comparison with MIBiG reference BGCs allowed assignment of 29 gene cluster families (GCFs) to pathways responsible for the production of known compounds, while for 57 GCFs, the molecular products remain unknown.
Authors: Koen Hoogendoorn, Lena Barra, Cees Waalwijk, Jeroen S. Dickschat, Theo A. J. van der Lee, Marnix H. Medema
Keywords: biosynthetic gene cluster, Fusarium, koraiol, ancestral state reconstruction (ASR), supernumary chromosome
Published: 05/06/2018
Repository: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5996196/
This work has received funding form the the EU’s H2020 research and innovation programme under GA No 678781- MycoKey.
CC-BY license http://creativecommons.org/